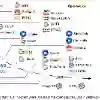
While the shift toward unified foundation models has revolutionized many deep learning domains, sleep medicine remains largely restricted to task-specific models that focus on localized micro-structure features. These approaches often neglect the rich, multi-modal context of Polysomnography (PSG) and fail to capture the global macro-structure of a full night's sleep. To address this, we introduce SleepMaMi , a Sleep Foundation Model engineered to master both hour-long sleep architectures and fine-grained signal morphologies. Our framework utilizes a hierarchical dual-encoder design: a Macro-Encoder to model full-night temporal dependencies and a Micro-Encoder to capture short-term characteristics from biosignals. Macro-Encoder is trained via Demographic-Guided Contrastive Learning, which aligns overnight sleep patterns with objective subject metadata, such as age, sex and BMI to refine global representations. Micro-Encoder is optimized via a hybrid Masked Autoencoder (MAE) and multi-modal contrastive objective. Pre-trained on a massive corpus of $>$20,000 PSG recordings (158K hours),SleepMaMi outperforms existing foundation models across a diverse suite of downstream tasks, demonstrating superior generalizability and label-efficient adaptation for clinical sleep analysis.
翻译:尽管向统一基础模型的转变已彻底革新了许多深度学习领域,睡眠医学在很大程度上仍局限于专注于局部微观结构特征的任务特定模型。这些方法通常忽略了多导睡眠图(PSG)丰富的多模态上下文,且未能捕捉整夜睡眠的全局宏观结构。为解决这一问题,我们提出了SleepMaMi,一种旨在掌握长达数小时的睡眠架构与细粒度信号形态的睡眠基础模型。我们的框架采用分层双编码器设计:宏观编码器用于建模整夜时间依赖性,微观编码器用于从生物信号中捕获短期特征。宏观编码器通过人口统计学引导的对比学习进行训练,该学习将整夜睡眠模式与客观受试者元数据(如年龄、性别和BMI)对齐,以优化全局表征。微观编码器通过混合掩码自编码器(MAE)和多模态对比目标进行优化。在超过20,000条PSG记录(158K小时)的大规模语料库上进行预训练后,SleepMaMi在一系列多样化的下游任务中超越了现有基础模型,展现了卓越的泛化能力和面向临床睡眠分析的标签高效适应性。