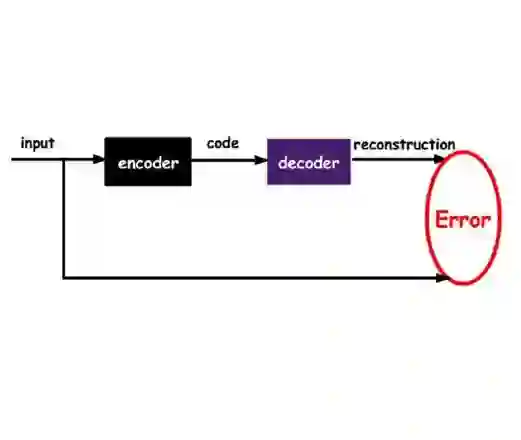
Many real-world problems require reasoning across multiple scales, demanding models which operate not on single data points, but on entire distributions. We introduce generative distribution embeddings (GDE), a framework that lifts autoencoders to the space of distributions. In GDEs, an encoder acts on sets of samples, and the decoder is replaced by a generator which aims to match the input distribution. This framework enables learning representations of distributions by coupling conditional generative models with encoder networks which satisfy a criterion we call distributional invariance. We show that GDEs learn predictive sufficient statistics embedded in the Wasserstein space, such that latent GDE distances approximately recover the $W_2$ distance, and latent interpolation approximately recovers optimal transport trajectories for Gaussian and Gaussian mixture distributions. We systematically benchmark GDEs against existing approaches on synthetic datasets, demonstrating consistently stronger performance. We then apply GDEs to six key problems in computational biology: learning donor-level representations from single-nuclei RNA sequencing data (6M cells), capturing clonal dynamics in lineage-traced RNA sequencing data (150K cells), predicting perturbation effects on transcriptomes (1M cells), predicting perturbation effects on cellular phenotypes (20M single-cell images), designing synthetic yeast promoters (34M sequences), and spatiotemporal modeling of viral protein sequences (1M sequences).
翻译:许多现实世界问题需要在多个尺度上进行推理,这要求模型不仅处理单个数据点,而且处理整个分布。我们提出生成式分布嵌入(GDE),一个将自编码器提升至分布空间的框架。在GDE中,编码器作用于样本集合,解码器被替换为旨在匹配输入分布的生成器。该框架通过将条件生成模型与满足我们称为分布不变性准则的编码器网络相结合,实现了对分布表征的学习。我们证明GDE学习嵌入在Wasserstein空间中的预测充分统计量,使得潜在GDE距离近似恢复$W_2$距离,且潜在插值近似恢复高斯分布和高斯混合分布的最优传输轨迹。我们在合成数据集上系统地将GDE与现有方法进行基准测试,证明其具有持续更强的性能。随后,我们将GDE应用于计算生物学中的六个关键问题:从单核RNA测序数据(600万细胞)中学习供体水平表征、在谱系追踪RNA测序数据(15万细胞)中捕捉克隆动态、预测扰动对转录组的影响(100万细胞)、预测扰动对细胞表型的影响(2000万单细胞图像)、设计合成酵母启动子(3400万序列)以及病毒蛋白序列的时空建模(100万序列)。