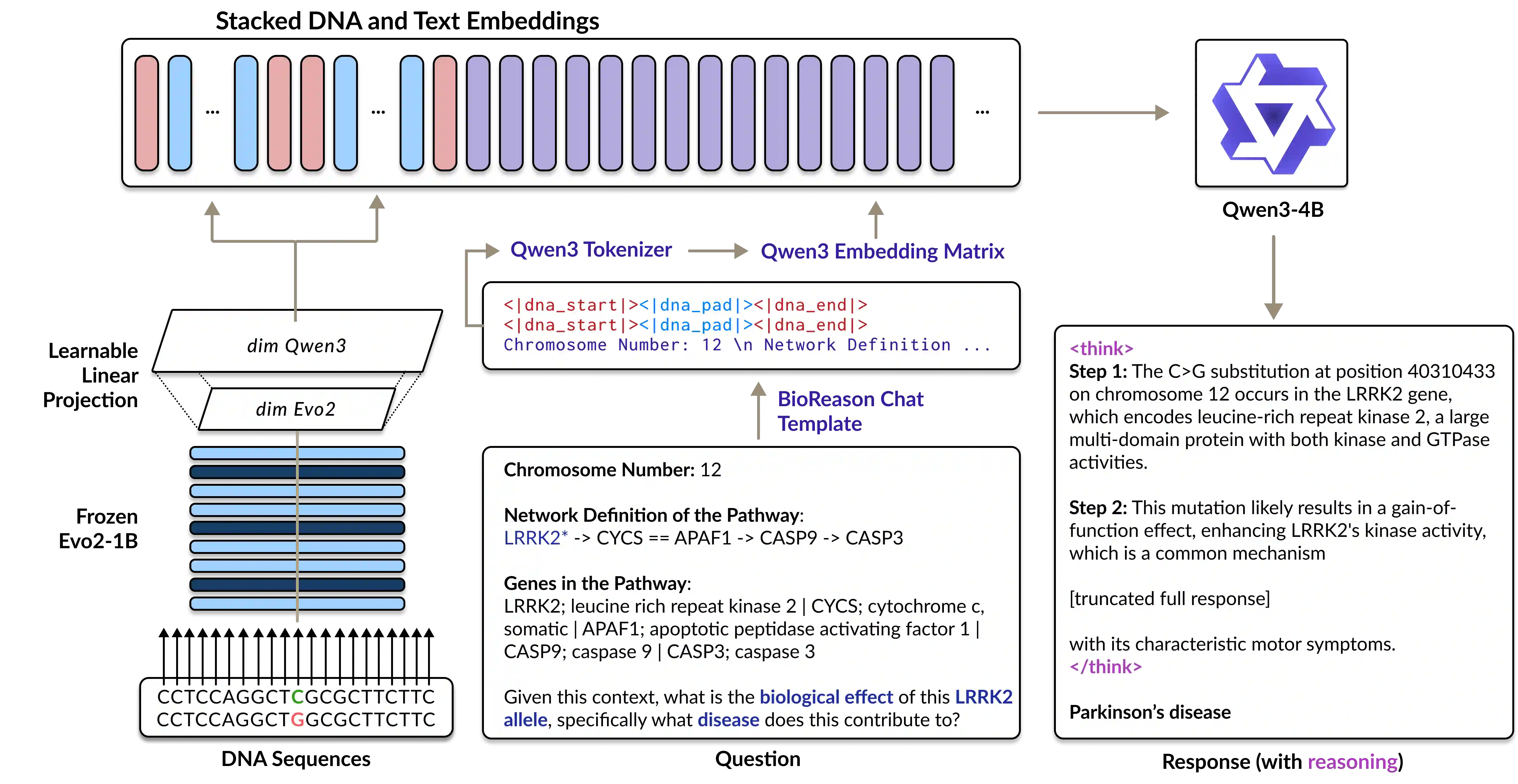
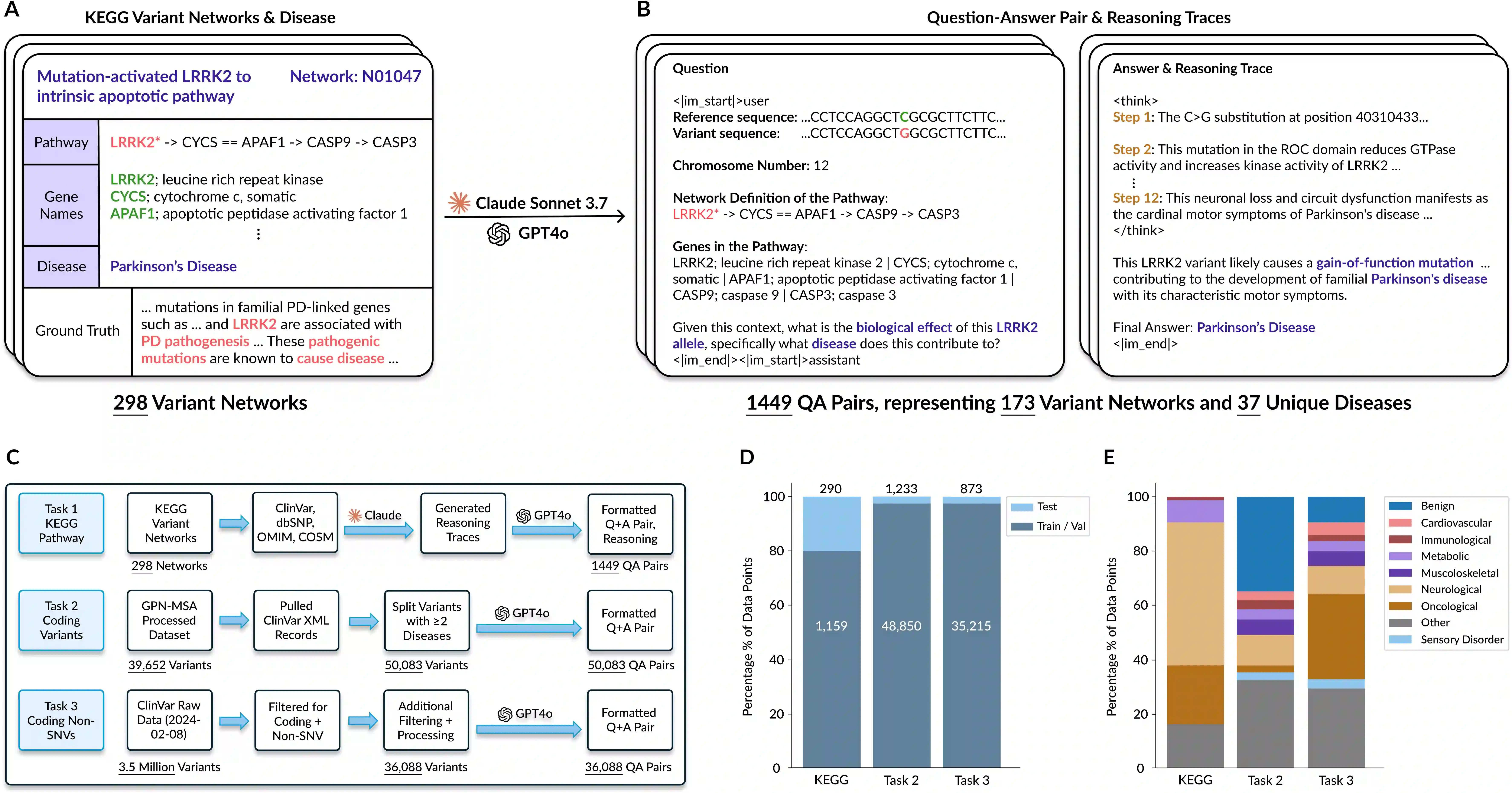
Unlocking deep and interpretable biological reasoning from complex genomic data remains a major AI challenge limiting scientific progress. While current DNA foundation models excel at representing sequences, they struggle with multi-step reasoning and lack transparent, biologically meaningful explanations. BioReason addresses this by tightly integrating a DNA foundation model with a large language model (LLM), enabling the LLM to directly interpret and reason over genomic information. Through supervised fine-tuning and reinforcement learning, BioReason learns to produce logical, biologically coherent deductions. It achieves major performance gains, boosting KEGG-based disease pathway prediction accuracy from 86% to 98% and improving variant effect prediction by an average of 15% over strong baselines. BioReason can reason over unseen biological entities and explain its decisions step by step, offering a transformative framework for interpretable, mechanistic AI in biology. All data, code, and checkpoints are available at https://github.com/bowang-lab/BioReason
翻译:从复杂的基因组数据中解锁深度且可解释的生物推理,仍然是限制科学进展的一项重大AI挑战。当前的DNA基础模型虽然在序列表征方面表现出色,但在多步推理方面存在困难,且缺乏透明、具有生物学意义的解释。BioReason通过将DNA基础模型与大型语言模型紧密集成来解决这一问题,使LLM能够直接解释基因组信息并基于其进行推理。通过监督微调和强化学习,BioReason学会生成逻辑严密、生物学上连贯的推论。它实现了显著的性能提升,将基于KEGG的疾病通路预测准确率从86%提高到98%,并将变异效应预测的平均性能在强基线基础上提升了15%。BioReason能够对未见过的生物实体进行推理,并逐步解释其决策过程,为生物学领域提供了一个变革性的、可解释的机制性AI框架。所有数据、代码和模型检查点均可在 https://github.com/bowang-lab/BioReason 获取。