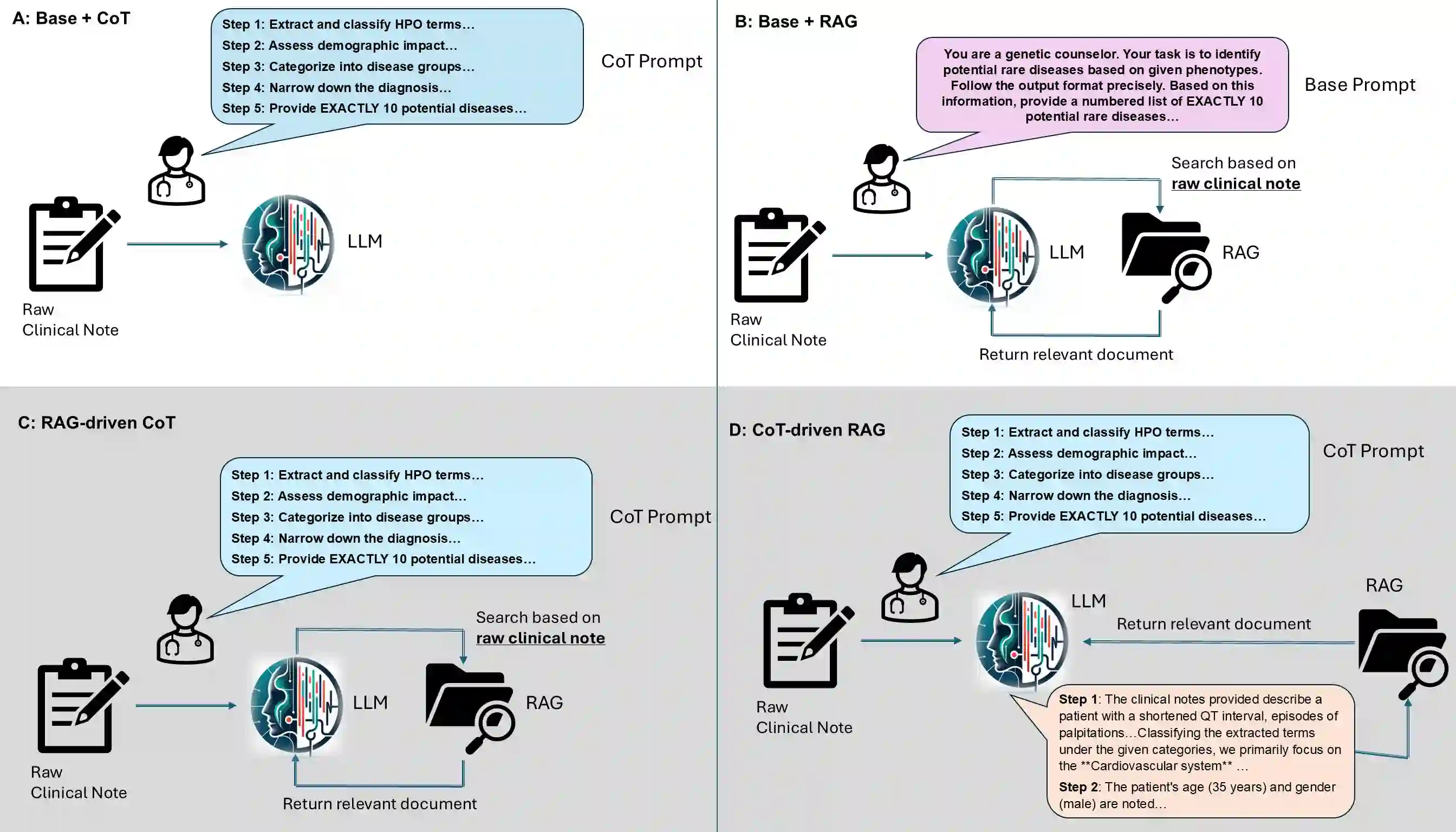
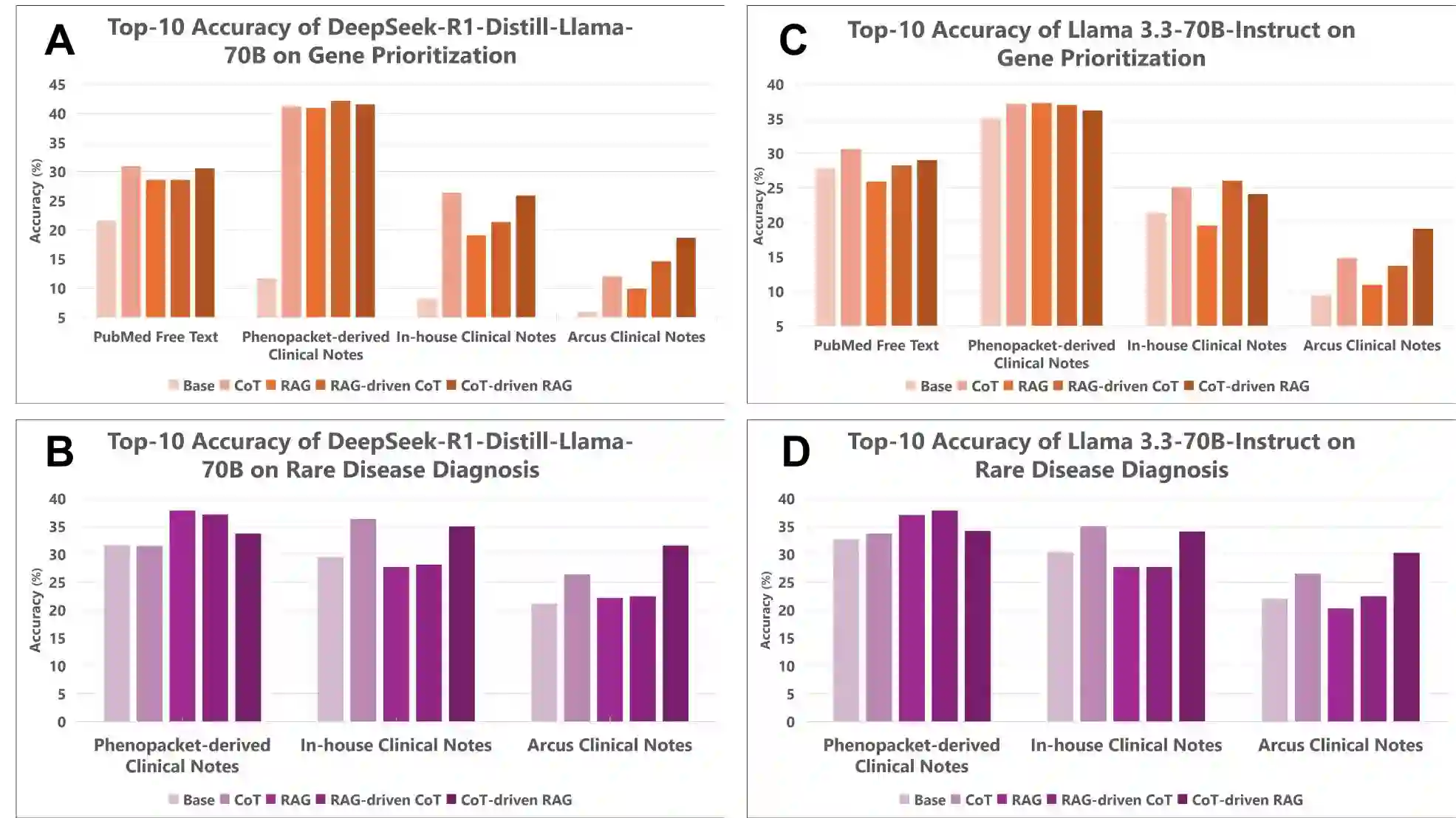
Background: Several studies show that large language models (LLMs) struggle with phenotype-driven gene prioritization for rare diseases. These studies typically use Human Phenotype Ontology (HPO) terms to prompt foundation models like GPT and LLaMA to predict candidate genes. However, in real-world settings, foundation models are not optimized for domain-specific tasks like clinical diagnosis, yet inputs are unstructured clinical notes rather than standardized terms. How LLMs can be instructed to predict candidate genes or disease diagnosis from unstructured clinical notes remains a major challenge. Methods: We introduce RAG-driven CoT and CoT-driven RAG, two methods that combine Chain-of-Thought (CoT) and Retrieval Augmented Generation (RAG) to analyze clinical notes. A five-question CoT protocol mimics expert reasoning, while RAG retrieves data from sources like HPO and OMIM (Online Mendelian Inheritance in Man). We evaluated these approaches on rare disease datasets, including 5,980 Phenopacket-derived notes, 255 literature-based narratives, and 220 in-house clinical notes from Childrens Hospital of Philadelphia. Results: We found that recent foundations models, including Llama 3.3-70B-Instruct and DeepSeek-R1-Distill-Llama-70B, outperformed earlier versions such as Llama 2 and GPT-3.5. We also showed that RAG-driven CoT and CoT-driven RAG both outperform foundation models in candidate gene prioritization from clinical notes; in particular, both methods with DeepSeek backbone resulted in a top-10 gene accuracy of over 40% on Phenopacket-derived clinical notes. RAG-driven CoT works better for high-quality notes, where early retrieval can anchor the subsequent reasoning steps in domain-specific evidence, while CoT-driven RAG has advantage when processing lengthy and noisy notes.
翻译:背景:多项研究表明,大型语言模型(LLM)在罕见病的表型驱动基因优先排序任务中表现欠佳。现有研究通常使用人类表型本体(HPO)术语提示GPT、LLaMA等基础模型以预测候选基因。然而在真实临床场景中,基础模型并未针对临床诊断等专业任务进行优化,且输入数据多为非结构化的临床记录而非标准化术语。如何指导LLM根据非结构化临床记录预测候选基因或疾病诊断仍是重大挑战。方法:我们提出了RAG驱动的CoT与CoT驱动的RAG两种方法,通过结合思维链(CoT)与检索增强生成(RAG)技术分析临床记录。其中五步提问式CoT协议模拟专家推理过程,RAG则从HPO、OMIM(在线人类孟德尔遗传数据库)等知识源检索数据。我们在罕见病数据集上评估了这些方法,包括5,980份基于Phenopacket生成的临床记录、255份文献叙述性病例及220份来自费城儿童医院的内部临床记录。结果:研究发现,包括Llama 3.3-70B-Instruct和DeepSeek-R1-Distill-Llama-70B在内的新版基础模型,其表现优于Llama 2、GPT-3.5等早期版本。实验同时证明,在基于临床记录的候选基因优先排序任务中,RAG驱动的CoT与CoT驱动的RAG均优于基础模型;特别值得注意的是,采用DeepSeek架构的两种方法在Phenopacket生成的临床记录上实现了超过40%的top-10基因准确率。对于高质量临床记录,RAG驱动的CoT表现更佳,其早期检索步骤能为后续推理提供领域证据锚点;而处理冗长且含噪声的记录时,CoT驱动的RAG更具优势。