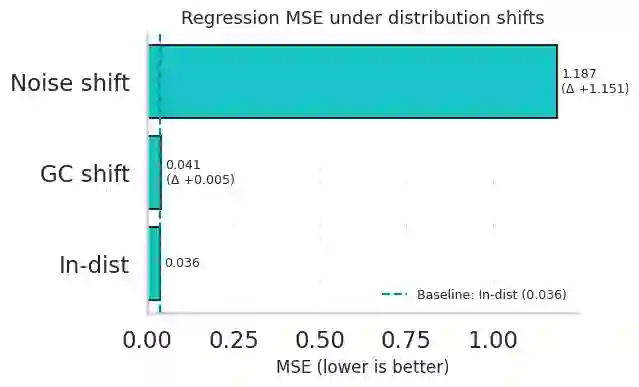
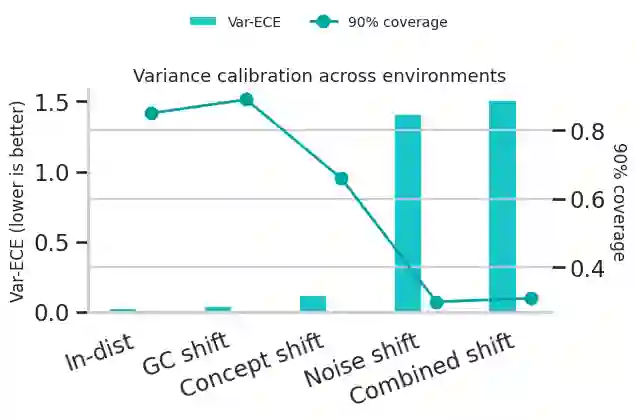
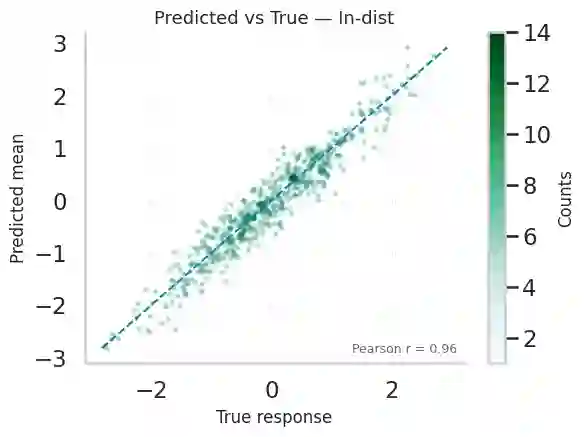
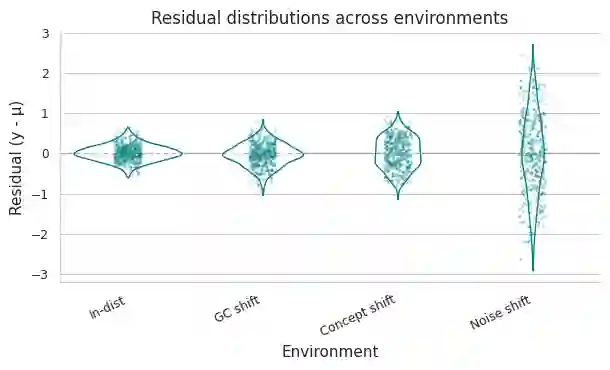
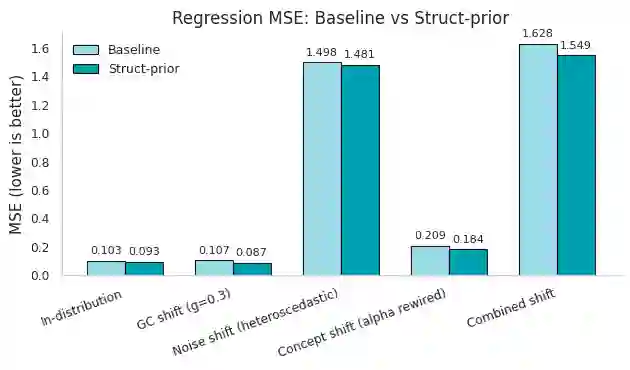
Robust machine learning for regulatory genomics is studied under biologically and technically induced distribution shifts. Deep convolutional and attention based models achieve strong in distribution performance on DNA regulatory sequence prediction tasks but are usually evaluated under i.i.d. assumptions, even though real applications involve cell type specific programs, evolutionary turnover, assay protocol changes, and sequencing artifacts. We introduce a robustness framework that combines a mechanistic simulation benchmark with real data analysis on a massively parallel reporter assay (MPRA) dataset to quantify performance degradation, calibration failures, and uncertainty based reliability. In simulation, motif driven regulatory outputs are generated with cell type specific programs, PWM perturbations, GC bias, depth variation, batch effects, and heteroscedastic noise, and CNN, BiLSTM, and transformer models are evaluated. Models remain accurate and reasonably calibrated under mild GC content shifts but show higher error, severe variance miscalibration, and coverage collapse under motif effect rewiring and noise dominated regimes, revealing robustness gaps invisible to standard i.i.d. evaluation. Adding simple biological structural priors motif derived features in simulation and global GC content in MPRA improves in distribution error and yields consistent robustness gains under biologically meaningful genomic shifts, while providing only limited protection against strong assay noise. Uncertainty-aware selective prediction offers an additional safety layer that risk coverage analyses on simulated and MPRA data show that filtering low confidence inputs recovers low risk subsets, including under GC-based out-of-distribution conditions, although reliability gains diminish when noise dominates.
翻译:本研究在生物和技术因素引起的分布偏移下,探讨调控基因组学中的鲁棒机器学习问题。深度卷积与基于注意力的模型在DNA调控序列预测任务中展现出优异的同分布性能,但现有评估通常基于独立同分布假设,而实际应用场景涉及细胞类型特异性调控程序、进化更替、检测协议变更及测序伪影等多种复杂因素。本文提出一个结合机制模拟基准与大规模并行报告基因检测(MPRA)数据集真实分析的鲁棒性评估框架,用于量化性能退化、校准失效及基于不确定性的可靠性指标。在模拟环境中,通过细胞类型特异性程序、PWM扰动、GC偏好、测序深度变异、批次效应和异方差噪声生成基序驱动的调控输出,并系统评估CNN、BiLSTM和Transformer模型的性能。研究发现:在温和的GC含量偏移下,模型能保持准确性与合理校准;但在基序效应重构和噪声主导机制下,模型呈现更高误差、严重的方差校准偏差及覆盖度崩溃,这些鲁棒性缺陷在标准独立同分布评估中无法显现。通过引入简单生物结构先验——在模拟中加入基序衍生特征,在MPRA数据中整合全局GC含量——可改善同分布误差,并在具有生物学意义的基因组偏移下获得一致的鲁棒性提升,但对强检测噪声的防护作用有限。不确定性感知的选择性预测提供了额外安全层:对模拟与MPRA数据的风险覆盖分析表明,过滤低置信度输入可恢复低风险子集(包括基于GC的分布外条件),但当噪声占主导时,可靠性增益显著减弱。