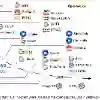
We present ImmuVis, an efficient convolutional foundation model for imaging mass cytometry (IMC), a high-throughput multiplex imaging technology that handles molecular marker measurements as image channels and enables large-scale spatial tissue profiling. Unlike natural images, multiplex imaging lacks a fixed channel space, as real-world marker sets vary across studies, violating a core assumption of standard vision backbones. To address this, ImmuVis introduces marker-adaptive hyperconvolutions that generate convolutional kernels from learned marker embeddings, enabling a single model to operate on arbitrary measured marker subsets without retraining. We pretrain ImmuVis on the largest to-date dataset, IMC17M (28 cohorts, 24,405 images, 265 markers, over 17M patches), using self-supervised masked reconstruction. ImmuVis outperforms SOTA baselines and ablations in virtual staining and downstream classification tasks at substantially lower compute cost than transformer-based alternatives, and is the sole model that provides calibrated uncertainty via a heteroscedastic likelihood objective. These results position ImmuVis as a practical, efficient foundation model for real-world IMC modeling.
翻译:本文提出ImmuVis,一种用于成像质谱流式细胞术(IMC)的高效卷积基础模型。IMC是一种高通量多重成像技术,将分子标记物测量值作为图像通道处理,实现大规模空间组织分析。与自然图像不同,多重成像缺乏固定的通道空间,因为实际研究中的标记物组合存在差异,这违背了标准视觉主干网络的核心假设。为解决此问题,ImmuVis引入标记物自适应超卷积,通过学习到的标记物嵌入生成卷积核,使单一模型能够处理任意测量的标记物子集而无需重新训练。我们在迄今最大数据集IMC17M(28个队列、24,405张图像、265种标记物、超过1700万个图像块)上使用自监督掩码重建任务对ImmuVis进行预训练。在虚拟染色和下游分类任务中,ImmuVis以显著低于基于Transformer的替代方案的算力成本超越了当前最先进的基线模型及消融实验,并且是唯一通过异方差似然目标提供校准不确定性的模型。这些成果使ImmuVis成为实际IMC建模中实用高效的基础模型。