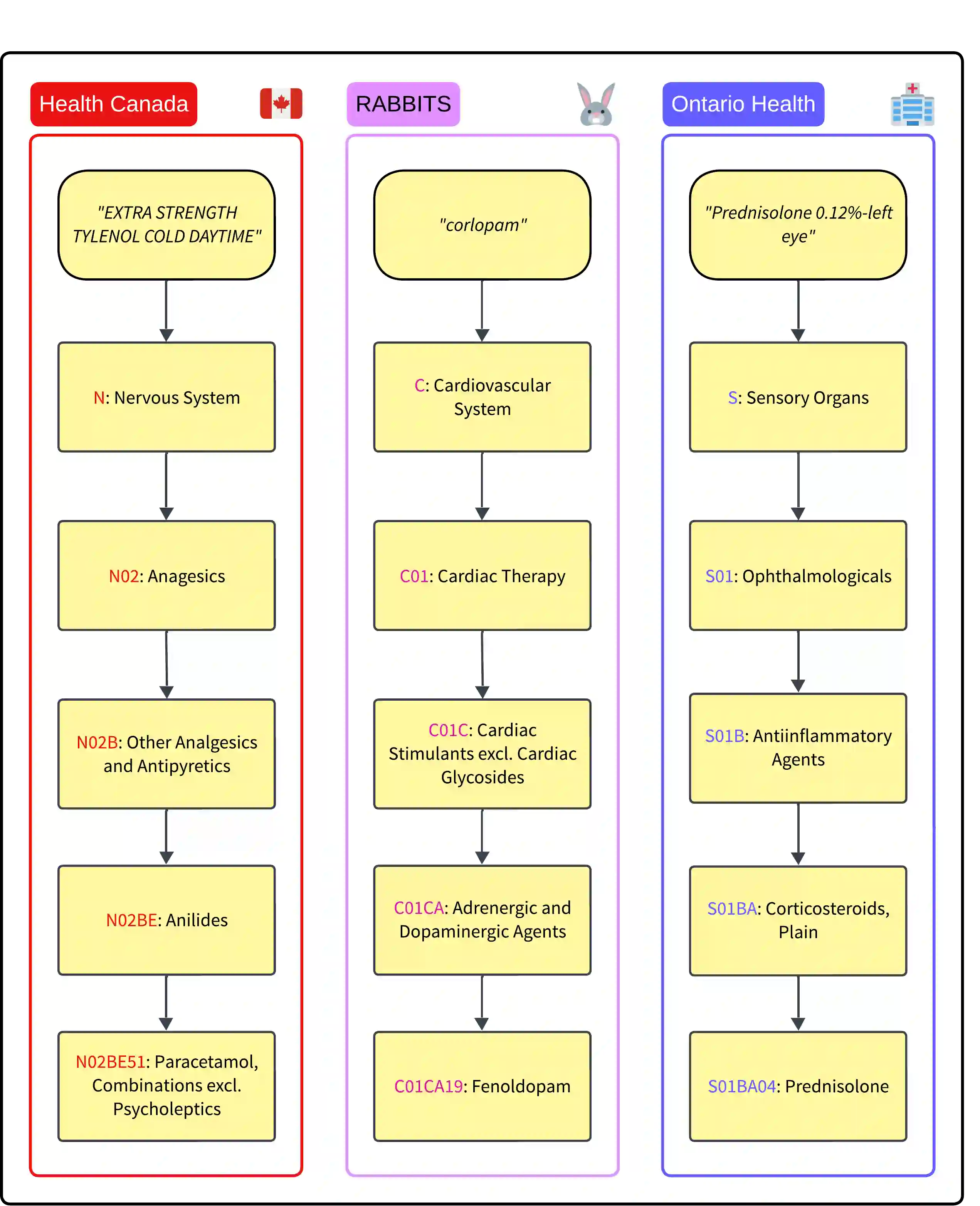
Manual assignment of Anatomical Therapeutic Chemical (ATC) codes to prescription records is a significant bottleneck in healthcare research and operations at Ontario Health and InterRAI Canada, requiring extensive expert time and effort. To automate this process while maintaining data privacy, we develop a practical approach using locally deployable large language models (LLMs). Inspired by recent advances in automatic International Classification of Diseases (ICD) coding, our method frames ATC coding as a hierarchical information extraction task, guiding LLMs through the ATC ontology level by level. We evaluate our approach using GPT-4o as an accuracy ceiling and focus development on open-source Llama models suitable for privacy-sensitive deployment. Testing across Health Canada drug product data, the RABBITS benchmark, and real clinical notes from Ontario Health, our method achieves 78% exact match accuracy with GPT-4o and 60% with Llama 3.1 70B. We investigate knowledge grounding through drug definitions, finding modest improvements in accuracy. Further, we show that fine-tuned Llama 3.1 8B matches zero-shot Llama 3.1 70B accuracy, suggesting that effective ATC coding is feasible with smaller models. Our results demonstrate the feasibility of automatic ATC coding in privacy-sensitive healthcare environments, providing a foundation for future deployments.
翻译:手动为处方记录分配解剖学治疗学化学分类法(ATC)编码是安大略省健康中心和InterRAI加拿大机构在医疗健康研究与运营中的一个显著瓶颈,需要耗费大量专家时间和精力。为了在保持数据隐私的同时实现该过程的自动化,我们开发了一种使用可本地部署的大语言模型(LLMs)的实用方法。受近期国际疾病分类(ICD)自动编码进展的启发,我们的方法将ATC编码构建为一个分层信息抽取任务,引导LLMs逐级遍历ATC本体。我们使用GPT-4o作为准确率上限来评估我们的方法,并将开发重点放在适合隐私敏感部署的开源Llama模型上。通过在加拿大卫生部药品数据、RABBITS基准测试以及来自安大略省健康中心的真实临床笔记上进行测试,我们的方法在GPT-4o上实现了78%的精确匹配准确率,在Llama 3.1 70B上实现了60%的准确率。我们研究了通过药物定义进行知识接地的效果,发现准确率有适度提升。此外,我们证明经过微调的Llama 3.1 8B模型能达到与零样本Llama 3.1 70B模型相当的准确率,这表明使用更小的模型实现有效的ATC编码是可行的。我们的结果证明了在隐私敏感的医疗健康环境中实现自动ATC编码的可行性,为未来的部署奠定了基础。