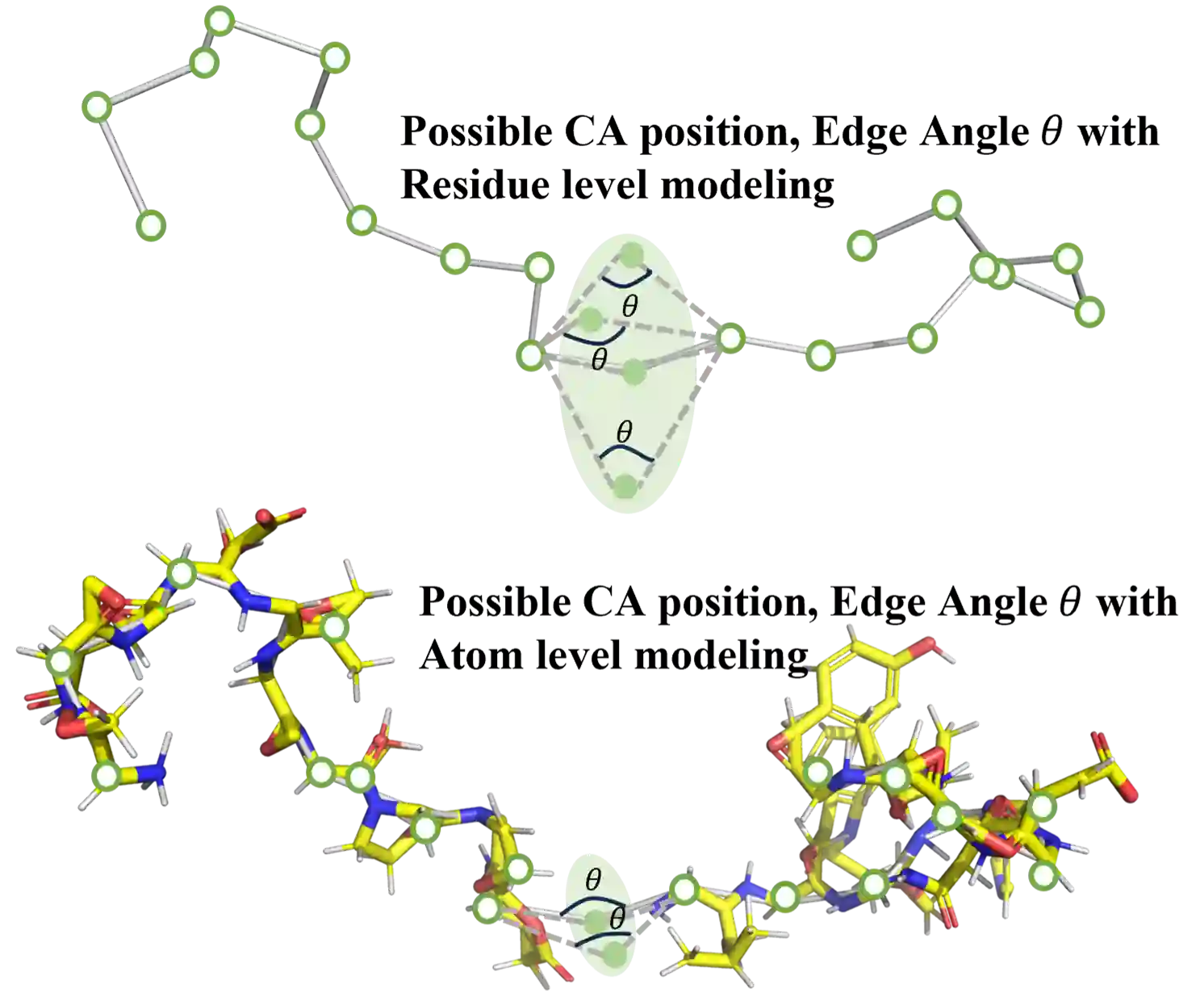
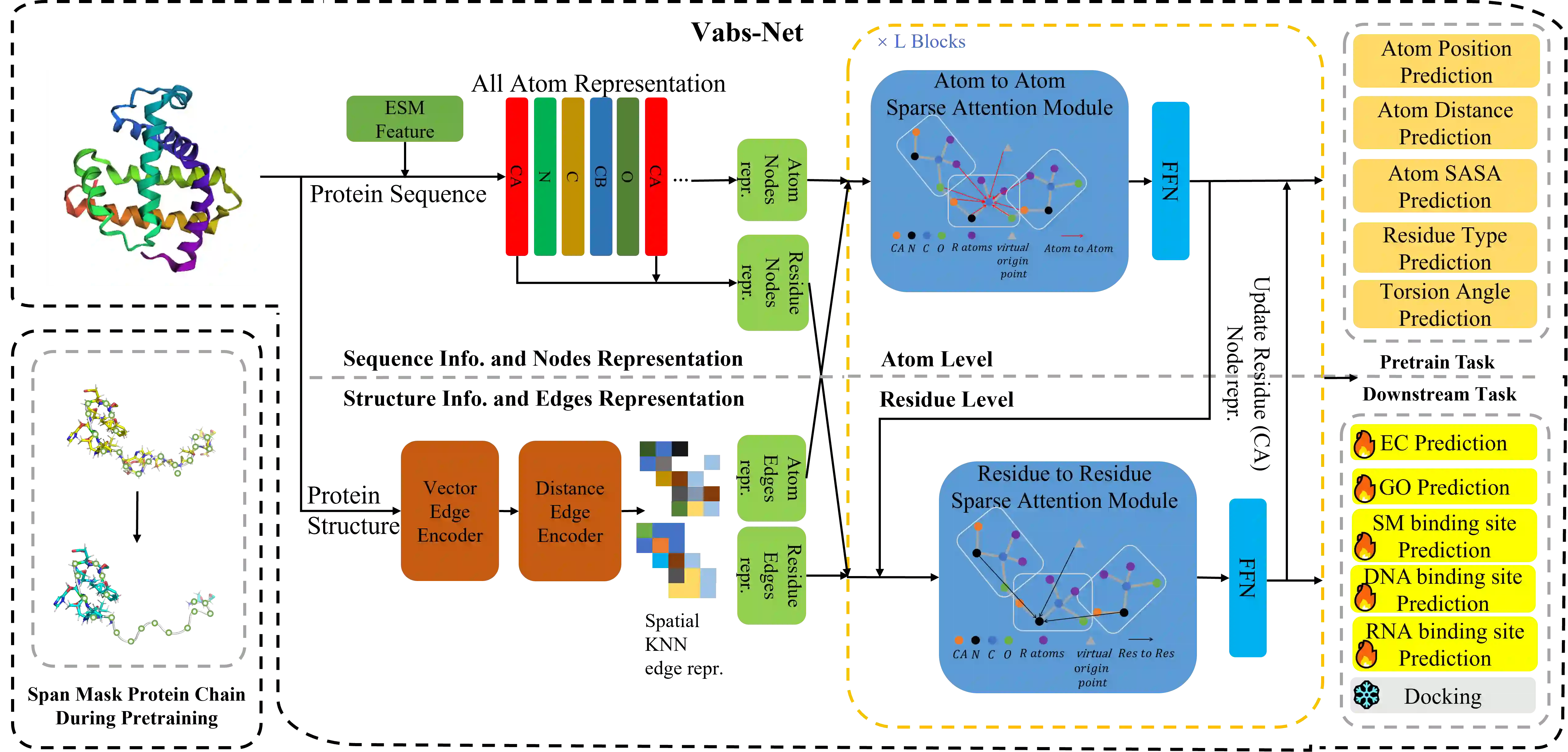
In recent years, there has been a surge in the development of 3D structure-based pre-trained protein models, representing a significant advancement over pre-trained protein language models in various downstream tasks. However, most existing structure-based pre-trained models primarily focus on the residue level, i.e., alpha carbon atoms, while ignoring other atoms like side chain atoms. We argue that modeling proteins at both residue and atom levels is important since the side chain atoms can also be crucial for numerous downstream tasks, for example, molecular docking. Nevertheless, we find that naively combining residue and atom information during pre-training typically fails. We identify a key reason is the information leakage caused by the inclusion of atom structure in the input, which renders residue-level pre-training tasks trivial and results in insufficiently expressive residue representations. To address this issue, we introduce a span mask pre-training strategy on 3D protein chains to learn meaningful representations of both residues and atoms. This leads to a simple yet effective approach to learning protein representation suitable for diverse downstream tasks. Extensive experimental results on binding site prediction and function prediction tasks demonstrate our proposed pre-training approach significantly outperforms other methods. Our code will be made public.
翻译:近年来,基于三维结构的预训练蛋白质模型发展迅猛,在各类下游任务中展现出相较于预训练蛋白质语言模型的显著优势。然而,现有基于结构的预训练模型主要聚焦于残基层级(即α碳原子),忽略了侧链原子等其他原子。我们认为,在残基和原子两个层级对蛋白质进行建模至关重要,因为侧链原子对分子对接等众多下游任务同样关键。但研究发现,在预训练过程中简单融合残基与原子信息通常会失败。我们识别出的关键原因在于:输入中包含原子结构会导致信息泄露,使残基层级的预训练任务变得过于简单,从而无法获得具有充分表达能力的残基表征。为解决这一问题,我们针对三维蛋白质链引入了跨度掩码预训练策略,以学习残基和原子的有意义的表征。由此形成了一种简单而有效的蛋白质表征学习方法,可适用于多种下游任务。在结合位点预测和功能预测任务上的大量实验结果表明,我们提出的预训练方法显著优于其他方法。我们将公开相关代码。