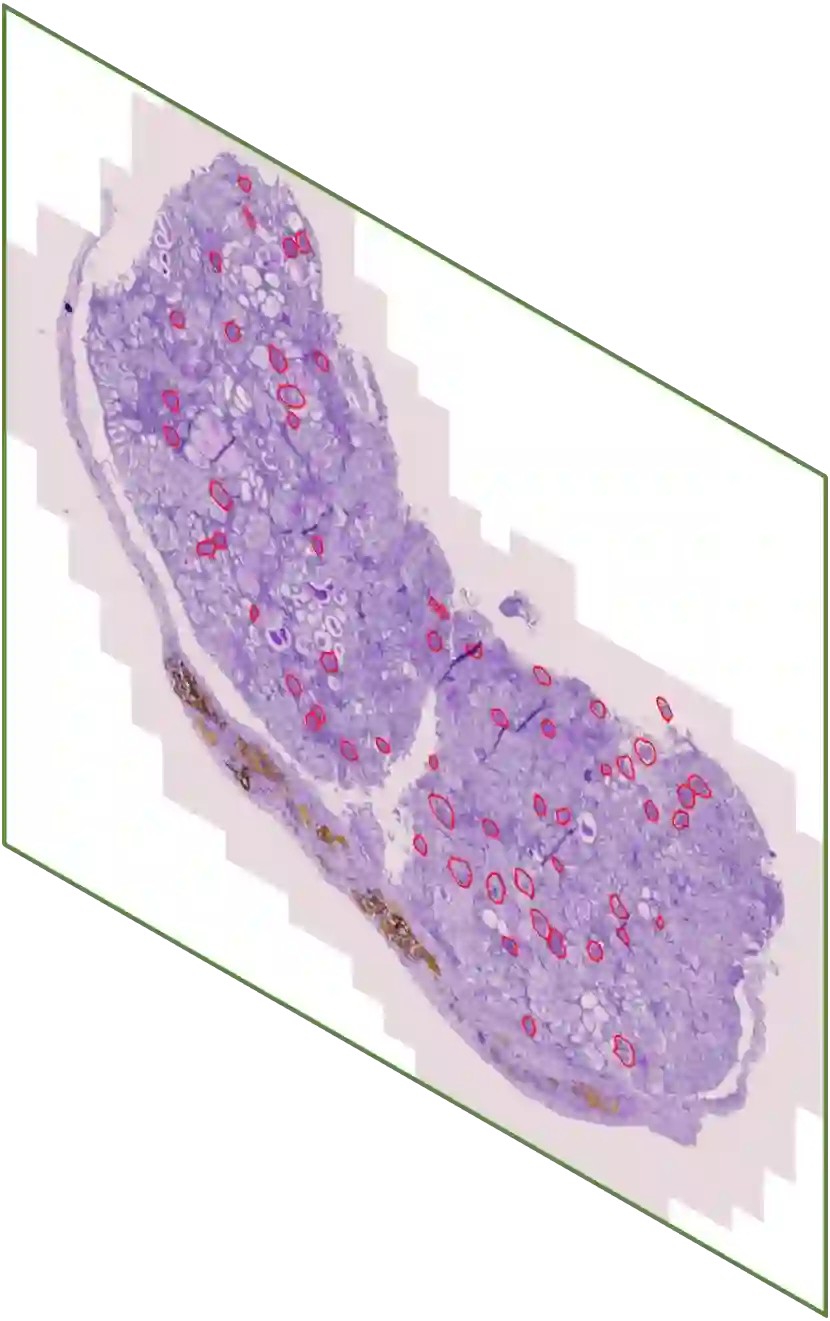
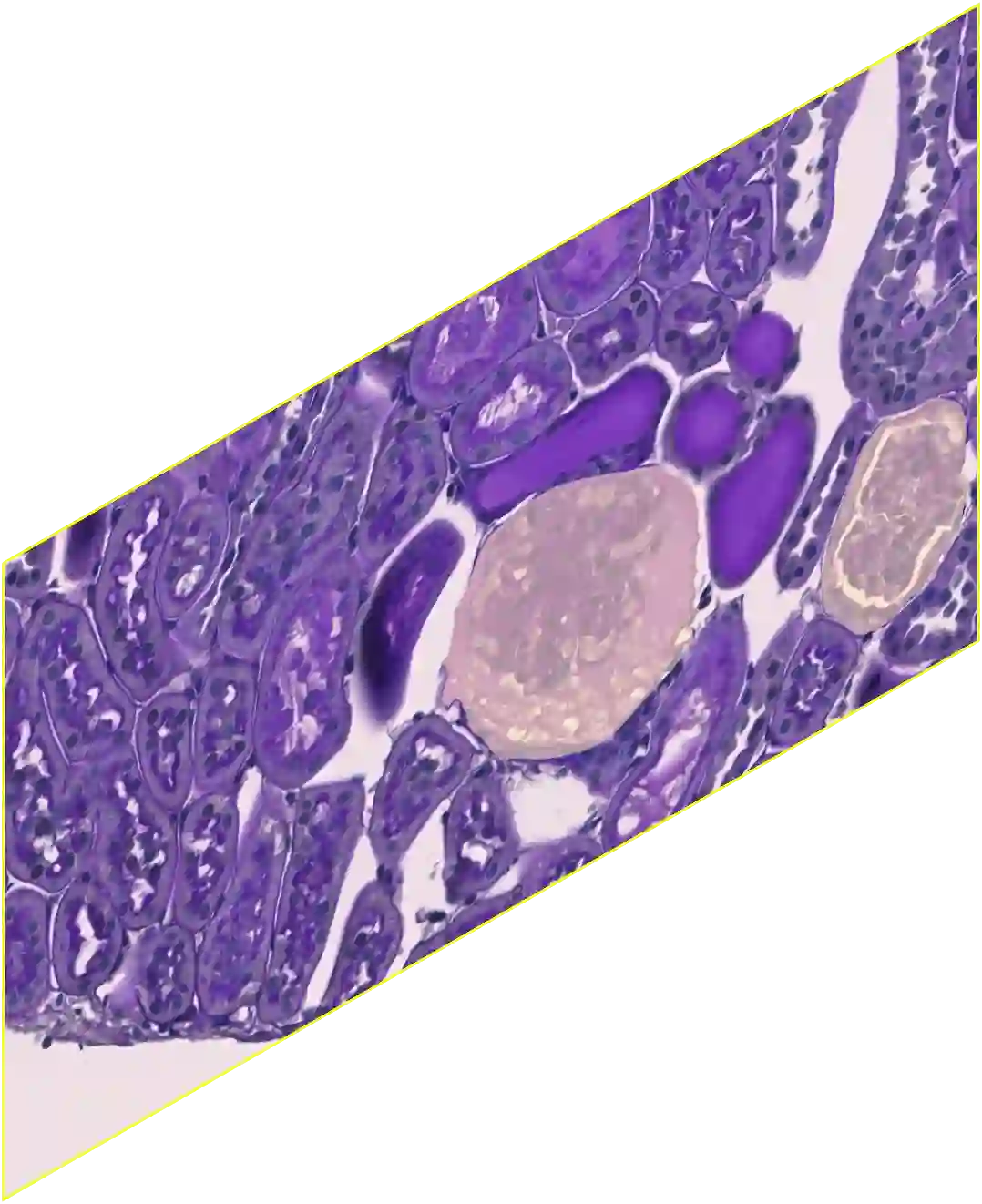
In digital pathology, the traditional method for deep learning-based image segmentation typically involves a two-stage process: initially segmenting high-resolution whole slide images (WSI) into smaller patches (e.g., 256x256, 512x512, 1024x1024) and subsequently reconstructing them to their original scale. This method often struggles to capture the complex details and vast scope of WSIs. In this paper, we propose the holistic histopathology (HoloHisto) segmentation method to achieve end-to-end segmentation on gigapixel WSIs, whose maximum resolution is above 80,000$\times$70,000 pixels. HoloHisto fundamentally shifts the paradigm of WSI segmentation to an end-to-end learning fashion with 1) a large (4K) resolution base patch for elevated visual information inclusion and efficient processing, and 2) a novel sequential tokenization mechanism to properly model the contextual relationships and efficiently model the rich information from the 4K input. To our best knowledge, HoloHisto presents the first holistic approach for gigapixel resolution WSI segmentation, supporting direct I/O of complete WSI and their corresponding gigapixel masks. Under the HoloHisto platform, we unveil a random 4K sampler that transcends ultra-high resolution, delivering 31 and 10 times more pixels than standard 2D and 3D patches, respectively, for advancing computational capabilities. To facilitate efficient 4K resolution dense prediction, we leverage sequential tokenization, utilizing a pre-trained image tokenizer to group image features into a discrete token grid. To assess the performance, our team curated a new kidney pathology image segmentation (KPIs) dataset with WSI-level glomeruli segmentation from whole mouse kidneys. From the results, HoloHisto-4K delivers remarkable performance gains over previous state-of-the-art models.
翻译:在数字病理学中,传统的基于深度学习的图像分割方法通常采用两阶段流程:首先将高分辨率全切片图像(WSI)分割为较小图块(例如256×256、512×512、1024×1024),随后将其重建至原始尺度。该方法往往难以有效捕捉全切片图像中复杂的细节信息与宏大的空间范围。本文提出整体组织病理学(HoloHisto)分割方法,首次实现对最大分辨率超过80,000×70,000像素的千兆像素级全切片图像进行端到端分割。HoloHisto通过以下两项创新从根本上改变了全切片图像分割的范式:1)采用大尺寸(4K分辨率)基础图块以提升视觉信息包容度与处理效率;2)引入新颖的序列标记化机制,有效建模上下文关联并高效处理4K输入中的丰富信息。据我们所知,HoloHisto是首个实现千兆像素分辨率全切片图像整体分割的方法,支持完整全切片图像及其对应千兆像素掩码的直接输入输出。在HoloHisto平台框架下,我们提出了超越超高分辨率限制的随机4K采样器,其提供的像素量分别达到标准2D与3D图块的31倍和10倍,显著提升了计算能力。为实现高效的4K分辨率密集预测,我们采用序列标记化技术,利用预训练图像标记器将图像特征聚合为离散标记网格。为评估方法性能,我们团队构建了新型肾脏病理图像分割(KPIs)数据集,包含全鼠肾脏的全切片图像级肾小球分割标注。实验结果表明,HoloHisto-4K相较于现有最优模型取得了显著的性能提升。