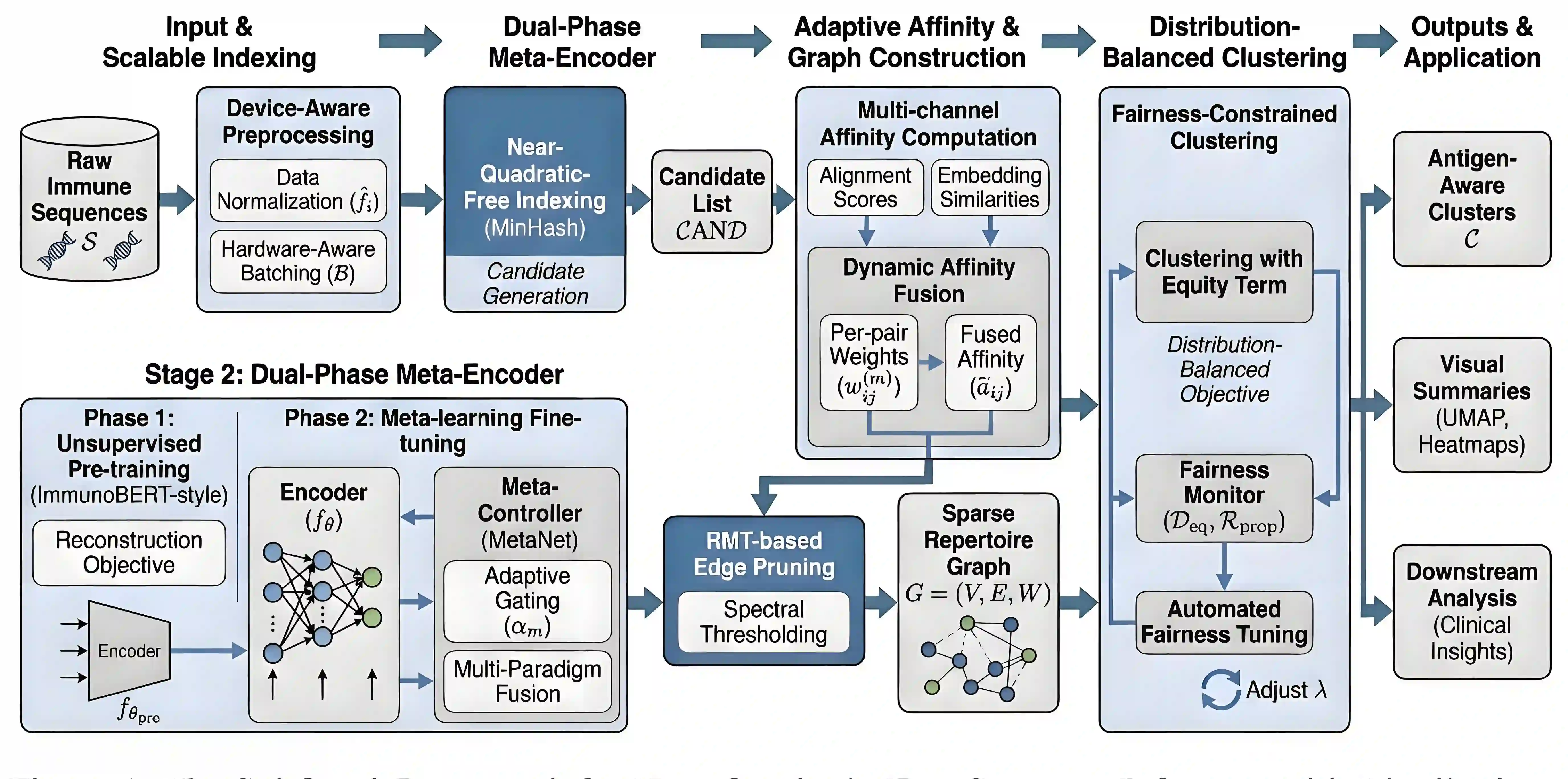
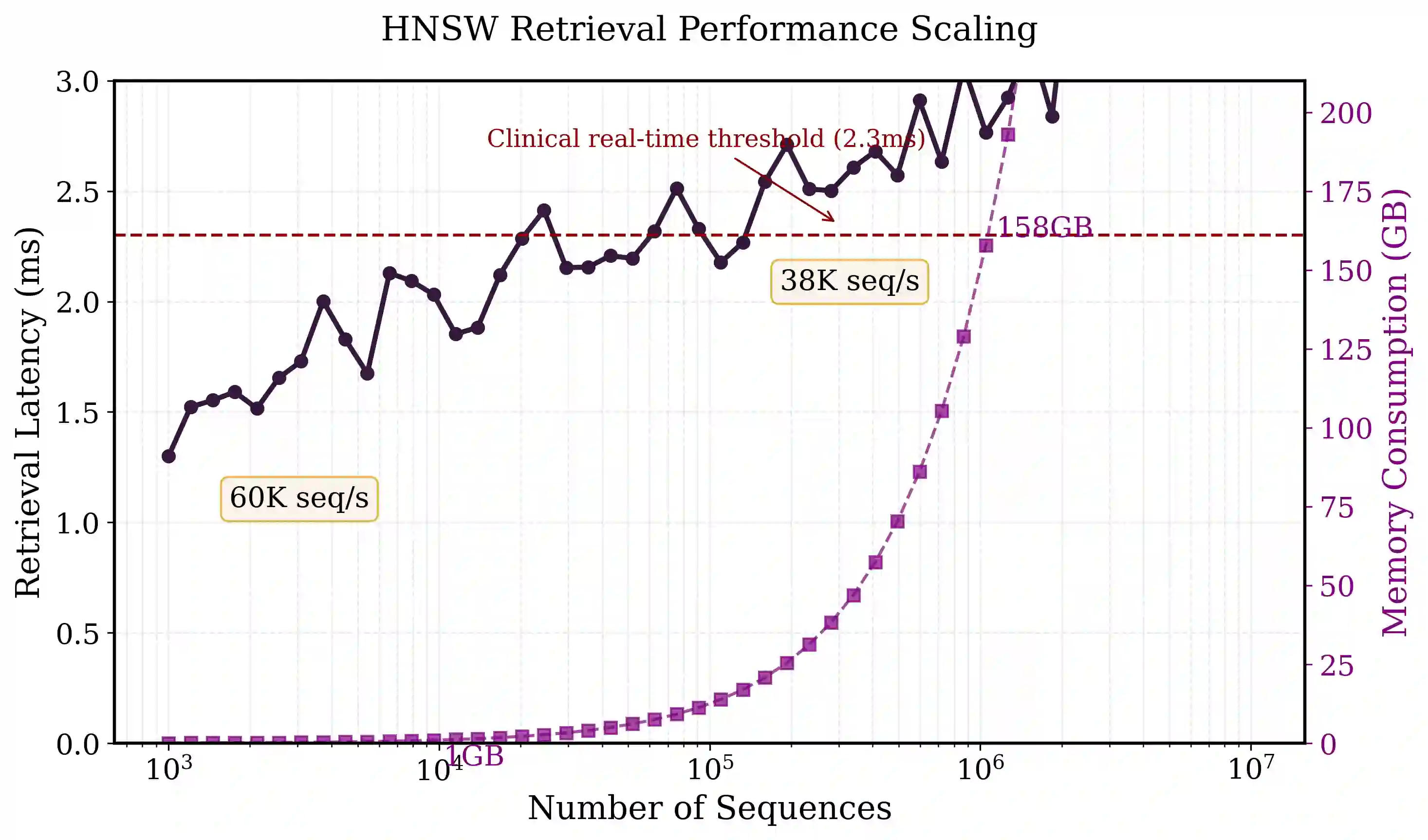
Comparative analysis of adaptive immune repertoires at population scale is hampered by two practical bottlenecks: the near-quadratic cost of pairwise affinity evaluations and dataset imbalances that obscure clinically important minority clonotypes. We introduce SubQuad, an end-to-end pipeline that addresses these challenges by combining antigen-aware, near-subquadratic retrieval with GPU-accelerated affinity kernels, learned multimodal fusion, and fairness-constrained clustering. The system employs compact MinHash prefiltering to sharply reduce candidate comparisons, a differentiable gating module that adaptively weights complementary alignment and embedding channels on a per-pair basis, and an automated calibration routine that enforces proportional representation of rare antigen-specific subgroups. On large viral and tumor repertoires SubQuad achieves measured gains in throughput and peak memory usage while preserving or improving recall@k, cluster purity, and subgroup equity. By co-designing indexing, similarity fusion, and equity-aware objectives, SubQuad offers a scalable, bias-aware platform for repertoire mining and downstream translational tasks such as vaccine target prioritization and biomarker discovery.
翻译:群体规模适应性免疫库的比较分析受到两个实际瓶颈的限制:成对亲和力评估的近二次方计算成本以及掩盖临床重要少数克隆型的数据集不平衡性。我们提出SubQuad,一种端到端流程,通过结合抗原感知的近次二次方检索、GPU加速的亲和力核函数、学习的多模态融合以及公平约束聚类来解决这些挑战。该系统采用紧凑的MinHash预过滤显著减少候选比较,通过可微分门控模块自适应地按对加权互补的比对和嵌入通道,并采用自动化校准程序确保稀有抗原特异性亚群的比例代表性。在大型病毒和肿瘤库上,SubQuad实现了吞吐量和峰值内存使用率的显著提升,同时保持或改进了召回率@k、聚类纯度及亚群公平性。通过协同设计索引、相似性融合和公平感知目标,SubQuad为库挖掘及下游转化任务(如疫苗靶点优先级排序和生物标志物发现)提供了一个可扩展的、偏差感知的平台。