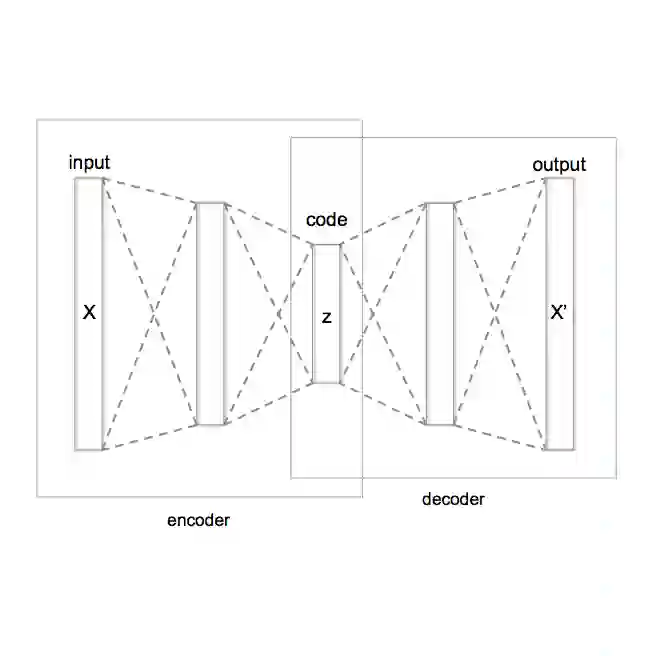
Generative models have emerged as powerful tools in medical imaging, enabling tasks such as segmentation, anomaly detection, and high-quality synthetic data generation. These models typically rely on learning meaningful latent representations, which are particularly valuable given the high-dimensional nature of 3D medical images like brain magnetic resonance imaging (MRI) scans. Despite their potential, latent representations remain underexplored in terms of their structure, information content, and applicability to downstream clinical tasks. Investigating these representations is crucial for advancing the use of generative models in neuroimaging research and clinical decision-making. In this work, we develop multiple variational autoencoders (VAEs) to encode 3D brain MRI scans into compact latent space representations for generative and predictive applications. We systematically evaluate the effectiveness of the learned representations through three key analyses: (i) a quantitative and qualitative assessment of MRI reconstruction quality, (ii) a visualisation of the latent space structure using Principal Component Analysis, and (iii) downstream classification tasks on a proprietary dataset of euploid and Down syndrome individuals brain MRI scans. Our results demonstrate that the VAE successfully captures essential brain features while maintaining high reconstruction fidelity. The latent space exhibits clear clustering patterns, particularly in distinguishing individuals with Down syndrome from euploid controls.
翻译:生成模型已成为医学影像领域的强大工具,能够实现分割、异常检测及高质量合成数据生成等任务。鉴于脑部磁共振成像(MRI)扫描等三维医学影像的高维特性,这些模型通常依赖于学习有意义的潜在表征,此类表征具有重要价值。尽管潜力巨大,但潜在表征在其结构、信息内容及对下游临床任务的适用性方面仍未得到充分探索。研究这些表征对于推动生成模型在神经影像学研究和临床决策中的应用至关重要。本研究开发了多种变分自编码器(VAEs),将三维脑部MRI扫描编码为紧凑的潜在空间表征,以用于生成与预测应用。我们通过三项关键分析系统评估了所学表征的有效性:(i)MRI重建质量的定量与定性评估,(ii)使用主成分分析对潜在空间结构的可视化呈现,以及(iii)在专有的整倍体与唐氏综合征个体脑部MRI扫描数据集上进行的下游分类任务。结果表明,VAE在保持高重建保真度的同时,成功捕获了关键的脑部特征。潜在空间展现出清晰的聚类模式,尤其在区分唐氏综合征个体与整倍体对照组方面表现显著。