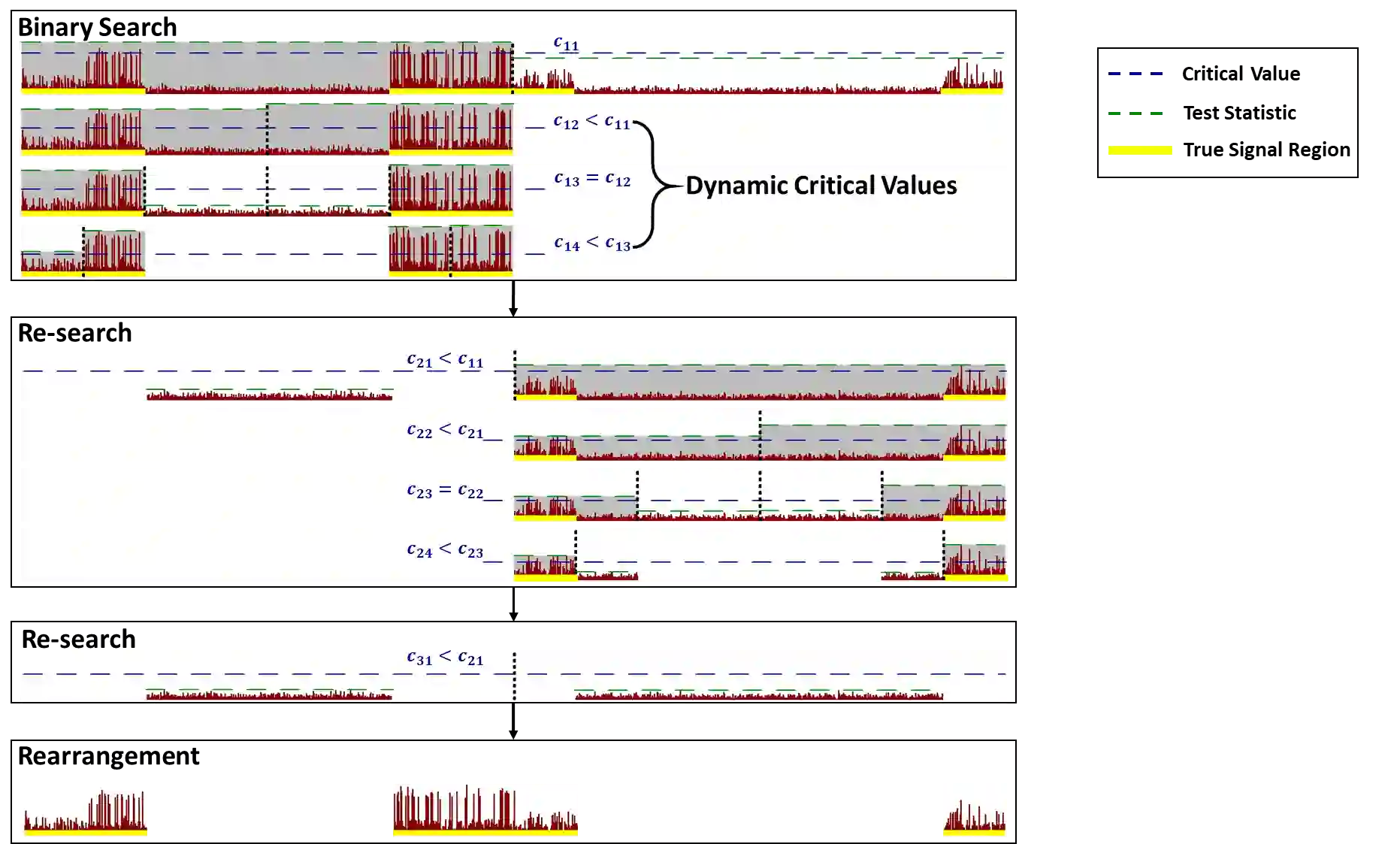
Research on the localization of the genetic basis associated with diseases or traits has been widely conducted in the last a few decades. Scan methods have been developed for region-based analysis in whole-genome association studies, helping us better understand how genetics influences human diseases or traits, especially when the aggregated effects of multiple causal variants are present. In this paper, we propose a fast and effective algorithm coupling with high-dimensional test for simultaneously detecting multiple signal regions, which is distinct from existing methods using scan or knockoff statistics. The idea is to conduct binary splitting with re-search and arrangement based on a sequence of dynamic critical values to increase detection accuracy and reduce computation. Theoretical and empirical studies demonstrate that our approach enjoys favorable theoretical guarantees with fewer restrictions and exhibits superior numerical performance with faster computation. Utilizing the UK Biobank data to identify the genetic regions related to breast cancer, we confirm previous findings and meanwhile, identify a number of new regions which suggest strong association with risk of breast cancer and deserve further investigation.
翻译:在过去的几十年中,关于疾病或性状相关遗传基础定位的研究已广泛开展。针对全基因组关联研究中的区域分析,扫描方法已被开发出来,帮助我们更好地理解遗传学如何影响人类疾病或性状,尤其是在存在多个因果变异聚合效应的情况下。本文提出了一种快速有效的算法,结合高维检验,用于同时检测多个信号区域,这与现有使用扫描或knockoff统计量的方法不同。其核心思想是基于一系列动态临界值进行带回溯与重排的二元分割,以提高检测精度并减少计算量。理论和实证研究表明,我们的方法具有更少限制的有利理论保证,并展现出更快的计算速度和优越的数值性能。利用英国生物银行数据识别与乳腺癌相关的遗传区域,我们确认了先前的发现,同时识别出许多新的区域,这些区域显示出与乳腺癌风险的强烈关联,值得进一步研究。