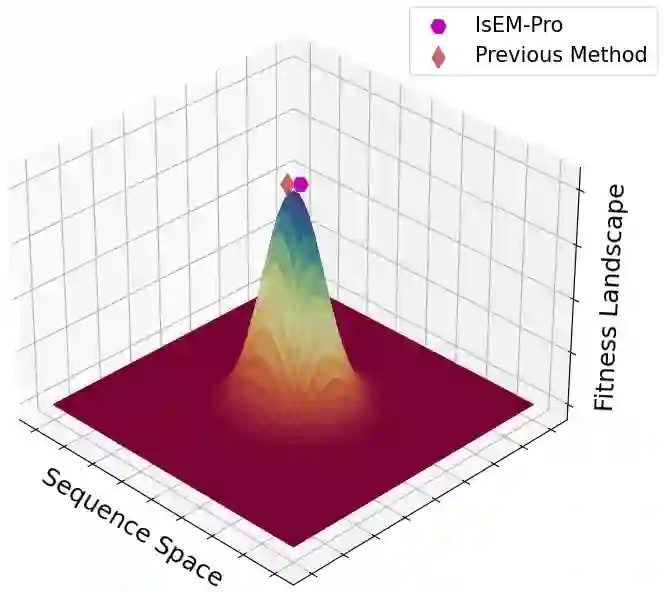
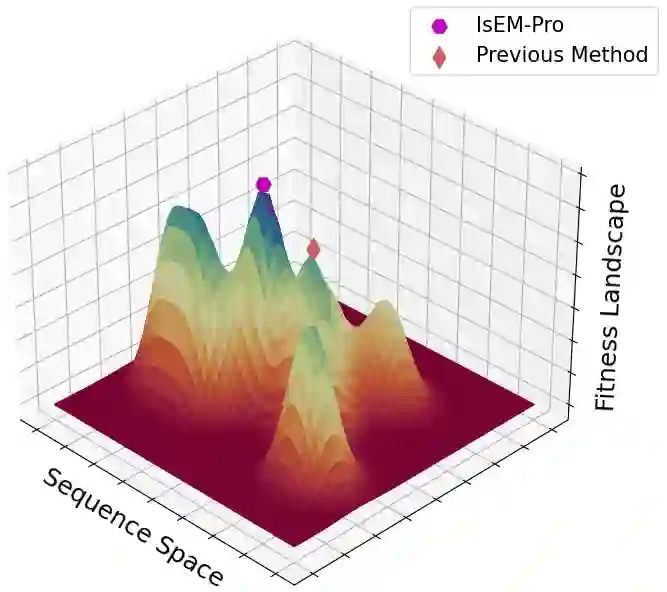
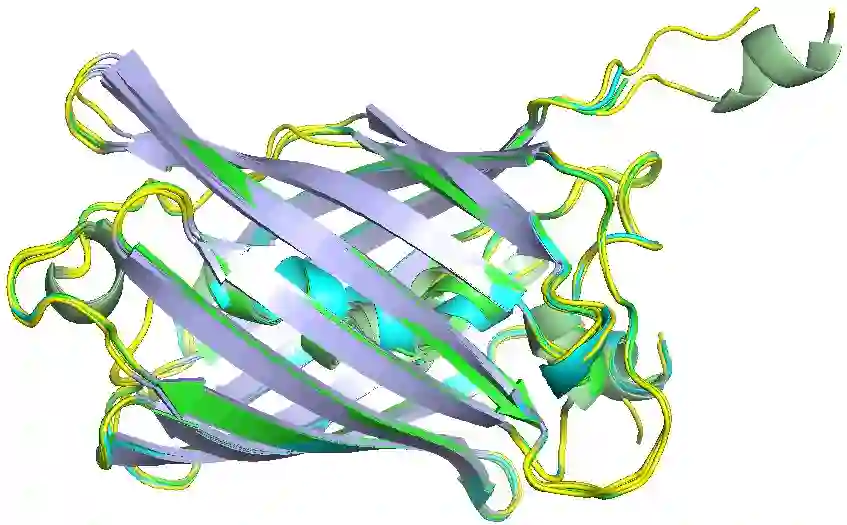
Designing protein sequences with desired biological function is crucial in biology and chemistry. Recent machine learning methods use a surrogate sequence-function model to replace the expensive wet-lab validation. How can we efficiently generate diverse and novel protein sequences with high fitness? In this paper, we propose IsEM-Pro, an approach to generate protein sequences towards a given fitness criterion. At its core, IsEM-Pro is a latent generative model, augmented by combinatorial structure features from a separately learned Markov random fields (MRFs). We develop an Monte Carlo Expectation-Maximization method (MCEM) to learn the model. During inference, sampling from its latent space enhances diversity while its MRFs features guide the exploration in high fitness regions. Experiments on eight protein sequence design tasks show that our IsEM-Pro outperforms the previous best methods by at least 55% on average fitness score and generates more diverse and novel protein sequences.
翻译:设计具有特定生物功能的蛋白质序列在生物学与化学领域至关重要。近期机器学习方法通过构建替代性序列-功能模型来替代昂贵的湿实验验证。如何高效生成具有高适应性的多样化新型蛋白质序列?本文提出IsEM-Pro方法,用于根据给定适应性标准生成蛋白质序列。该方法核心为潜在生成模型,并通过从独立学习的马尔可夫随机场中提取组合结构特征进行增强。我们开发了蒙特卡洛期望最大化算法来训练该模型。在推理过程中,从潜在空间采样可增强序列多样性,而马尔可夫随机场特征则引导模型在高适应性区域进行探索。在八个蛋白质序列设计任务上的实验表明,IsEM-Pro在平均适应性得分上较先前最优方法提升至少55%,并能生成更具多样性和新颖性的蛋白质序列。