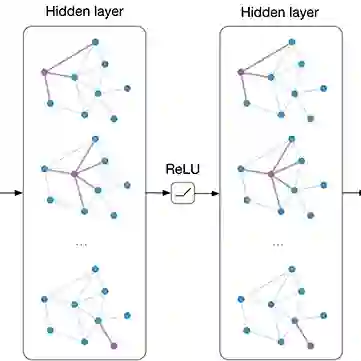
The extraction of chemical-gene relations plays a pivotal role in understanding the intricate interactions between chemical compounds and genes, with significant implications for drug discovery, disease understanding, and biomedical research. This paper presents a data set created by merging the ChemProt and DrugProt datasets to augment sample counts and improve model accuracy. We evaluate the merged dataset using two state of the art relationship extraction algorithms: Bidirectional Encoder Representations from Transformers (BERT) specifically BioBERT, and Graph Convolutional Networks (GCNs) combined with BioBERT. While BioBERT excels at capturing local contexts, it may benefit from incorporating global information essential for understanding chemical-gene interactions. This can be achieved by integrating GCNs with BioBERT to harness both global and local context. Our results show that by integrating the ChemProt and DrugProt datasets, we demonstrated significant improvements in model performance, particularly in CPR groups shared between the datasets. Incorporating the global context using GCN can help increase the overall precision and recall in some of the CPR groups over using just BioBERT.
翻译:化学-基因关系的抽取对于理解化学化合物与基因之间复杂的相互作用至关重要,对药物发现、疾病理解和生物医学研究具有重大意义。本文提出了通过合并ChemProt和DrugProt数据集构建的数据集,以增加样本数量并提升模型准确性。我们使用两种先进的关系抽取算法对合并数据集进行评估:基于Transformer的双向编码器表示(BERT),具体为BioBERT,以及图卷积网络(GCN)与BioBERT的结合。虽然BioBERT擅长捕捉局部上下文,但结合理解化学-基因相互作用所必需的全局信息可能使其受益。这可以通过将GCN与BioBERT集成来实现,从而同时利用全局和局部上下文。我们的结果表明,通过整合ChemProt和DrugProt数据集,模型性能得到显著提升,尤其是在两个数据集共享的CPR组中。使用GCN引入全局上下文有助于在某些CPR组中提高整体精确率和召回率,优于仅使用BioBERT的效果。