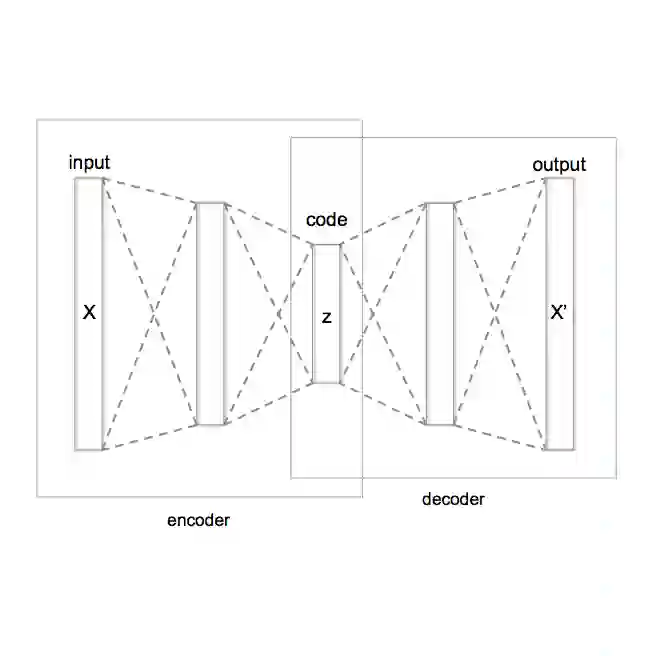
High-dimensional neuroimaging data presents challenges for assessing neurodegenerative diseases due to complex non-linear relationships. Variational Autoencoders (VAEs) can encode scans into lower-dimensional latent spaces capturing disease-relevant features. We propose a semi-supervised VAE framework with a flexible similarity regularization term that aligns selected latent variables with clinical or biomarker measures of dementia progression. This allows adapting the similarity metric and supervised variables to specific goals or available data. We demonstrate the approach using PET scans from the Alzheimer's Disease Neuroimaging Initiative (ADNI), guiding the first latent dimension to align with a cognitive score. Using this supervised latent variable, we generate average reconstructions across levels of cognitive impairment. Voxel-wise GLM analysis reveals reduced metabolism in key regions, mainly the hippocampus, and within major Resting State Networks, particularly the Default Mode and Central Executive Networks. The remaining latent variables encode affine transformations and intensity variations, capturing confounds such as inter-subject variability and site effects. Our framework effectively extracts disease-related patterns aligned with established Alzheimer's biomarkers, offering an interpretable and adaptable tool for studying neurodegenerative progression.
翻译:高维神经影像数据因其复杂的非线性关系,给评估神经退行性疾病带来了挑战。变分自编码器(VAEs)能够将扫描图像编码为低维潜在空间,以捕捉疾病相关特征。我们提出一种半监督VAE框架,其采用灵活的相似性正则化项,使选定的潜在变量与痴呆进展的临床或生物标志物指标对齐。这允许根据特定目标或可用数据调整相似性度量和监督变量。我们利用阿尔茨海默病神经影像倡议(ADNI)的PET扫描数据验证该方法,引导第一个潜在维度与认知评分对齐。使用这一监督潜在变量,我们生成了不同认知损伤水平下的平均重建图像。基于体素的广义线性模型分析揭示了关键区域(主要是海马体)以及主要静息态网络(特别是默认模式网络和中央执行网络)内的代谢降低。其余潜在变量编码仿射变换和强度变化,捕捉了如受试者间变异性和扫描站点效应等混杂因素。我们的框架有效提取了与已确立的阿尔茨海默病生物标志物一致的疾病相关模式,为研究神经退行性进展提供了一种可解释且适应性强的工具。