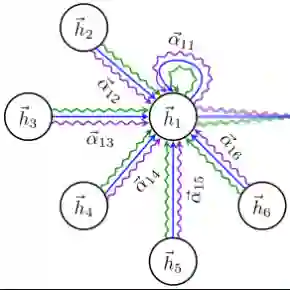
Protein-protein interactions (PPIs) play key roles in a broad range of biological processes. Numerous strategies have been proposed for predicting PPIs, and among them, graph-based methods have demonstrated promising outcomes owing to the inherent graph structure of PPI networks. This paper reviews various graph-based methodologies, and discusses their applications in PPI prediction. We classify these approaches into two primary groups based on their model structures. The first category employs Graph Neural Networks (GNN) or Graph Convolutional Networks (GCN), while the second category utilizes Graph Attention Networks (GAT), Graph Auto-Encoders and Graph-BERT. We highlight the distinctive methodologies of each approach in managing the graph-structured data inherent in PPI networks and anticipate future research directions in this domain.
翻译:蛋白质相互作用在广泛的生物过程中扮演关键角色。为预测蛋白质相互作用,研究者已提出众多策略,其中基于图的方法因蛋白质相互作用网络固有的图结构而展现出显著成效。本文综述了各类基于图的方法,并探讨其在蛋白质相互作用预测中的应用。根据模型结构,我们将这些方法分为两大类:第一类采用图神经网络或图卷积网络,第二类运用图注意力网络、图自编码器与Graph-BERT。本文着重阐述了各类方法在处理蛋白质相互作用网络图结构数据中的独特策略,并展望了该领域的未来研究方向。