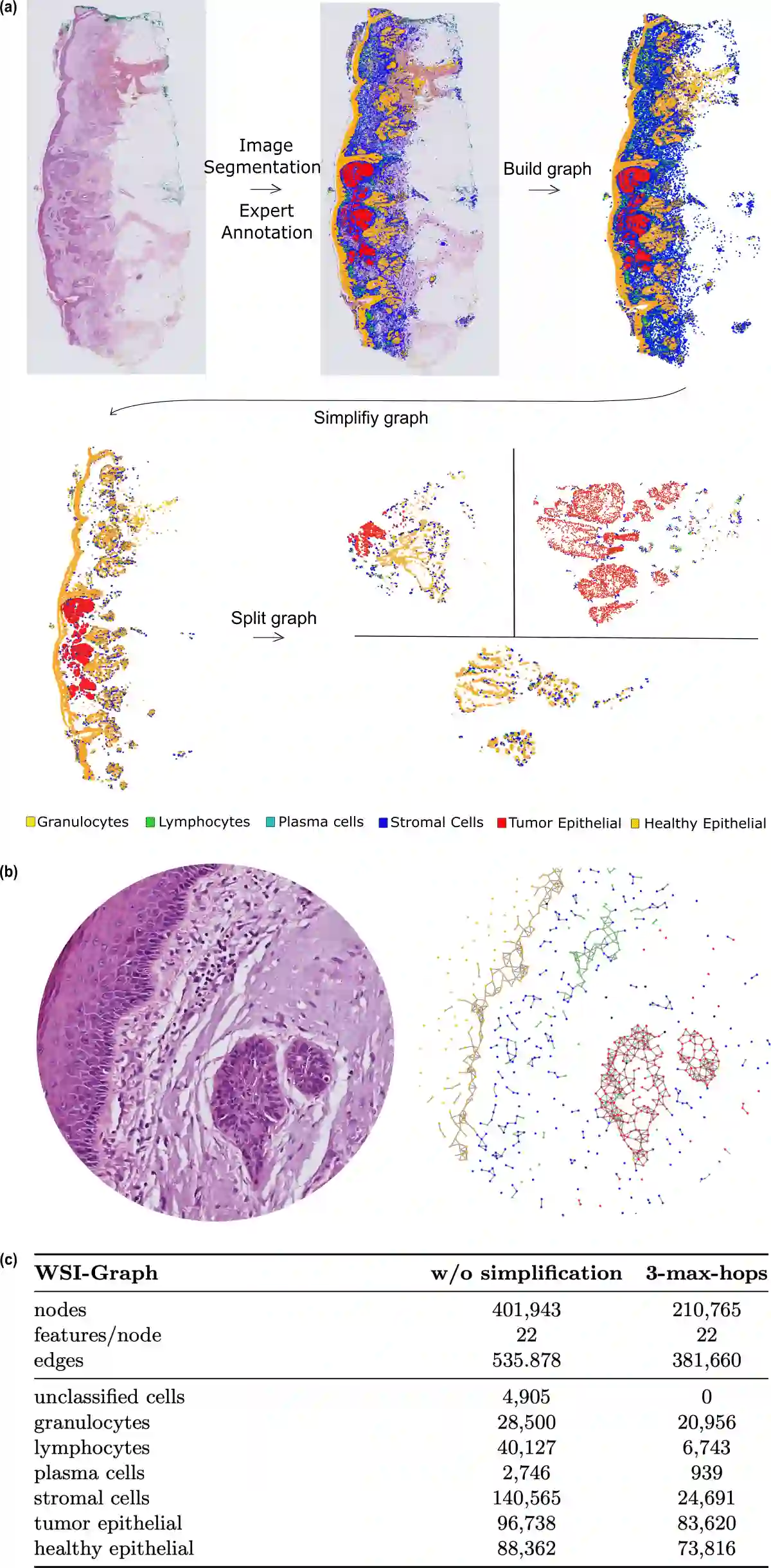
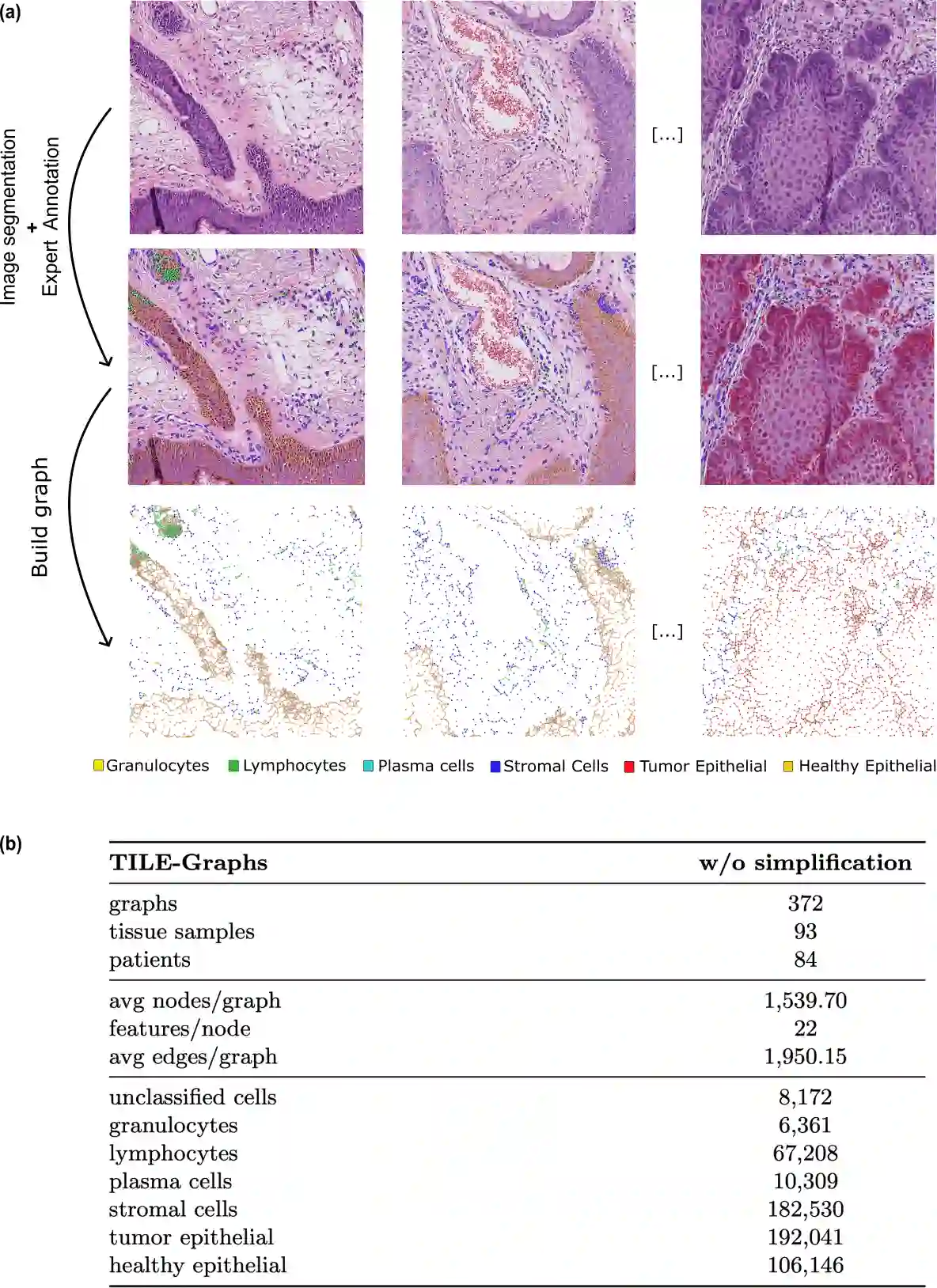
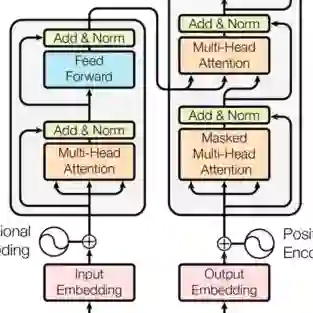
Whole-slide images (WSIs) from cancer patients contain rich information that can be used for medical diagnosis or to follow treatment progress. To automate their analysis, numerous deep learning methods based on convolutional neural networks and Vision Transformers have been developed and have achieved strong performance in segmentation and classification tasks. However, due to the large size and complex cellular organization of WSIs, these models rely on patch-based representations, losing vital tissue-level context. We propose using scalable Graph Transformers on a full-WSI cell graph for classification. We evaluate this methodology on a challenging task: the classification of healthy versus tumor epithelial cells in cutaneous squamous cell carcinoma (cSCC), where both cell types exhibit very similar morphologies and are therefore difficult to differentiate for image-based approaches. We first compared image-based and graph-based methods on a single WSI. Graph Transformer models SGFormer and DIFFormer achieved balanced accuracies of $85.2 \pm 1.5$ ($\pm$ standard error) and $85.1 \pm 2.5$ in 3-fold cross-validation, respectively, whereas the best image-based method reached $81.2 \pm 3.0$. By evaluating several node feature configurations, we found that the most informative representation combined morphological and texture features as well as the cell classes of non-epithelial cells, highlighting the importance of the surrounding cellular context. We then extended our work to train on several WSIs from several patients. To address the computational constraints of image-based models, we extracted four $2560 \times 2560$ pixel patches from each image and converted them into graphs. In this setting, DIFFormer achieved a balanced accuracy of $83.6 \pm 1.9$ (3-fold cross-validation), while the state-of-the-art image-based model CellViT256 reached $78.1 \pm 0.5$.
翻译:癌症患者的全切片图像包含丰富信息,可用于医学诊断或追踪治疗进展。为实现自动化分析,基于卷积神经网络和视觉Transformer的多种深度学习方法已被开发出来,并在分割与分类任务中取得优异性能。然而,由于全切片图像尺寸庞大且细胞组织结构复杂,这些模型依赖基于图像块的表示方法,丢失了关键的组织层面上下文信息。我们提出在全切片细胞图上使用可扩展图Transformer进行分类。我们在具有挑战性的任务上评估该方法:皮肤鳞状细胞癌中健康与肿瘤上皮细胞的分类,其中两种细胞类型形态极为相似,因此基于图像的方法难以区分。我们首先在单张全切片图像上比较了基于图像与基于图的方法。图Transformer模型SGFormer和DIFFormer在3折交叉验证中分别达到$85.2 \pm 1.5$($\pm$标准误)和$85.1 \pm 2.5$的平衡准确率,而最佳基于图像的方法为$81.2 \pm 3.0$。通过评估多种节点特征配置,我们发现最具信息量的表征结合了形态学特征、纹理特征以及非上皮细胞的细胞类别,凸显了周围细胞环境的重要性。随后我们将研究扩展至基于多患者的多张全切片图像进行训练。为应对基于图像模型的计算限制,我们从每张图像提取四个$2560 \times 2560$像素的图像块并将其转换为图结构。在此设置下,DIFFormer获得$83.6 \pm 1.9$的平衡准确率(3折交叉验证),而基于图像的先进模型CellViT256为$78.1 \pm 0.5$。